We sent a bone sample for DNA testing to the McMaster Ancient DNA Centre, McMaster University, Hamilton, Ontario two weeks ago. Andy Clack, a PhD student in the Centre, sent this encouraging reply:
I have the talus/ankle bone in the lab now… wow! That thing is like a rock. I couldn’t ask for a better specimen! I’m going to use a sterile dremel tool and a particular type of bit that generates less heat and produces shards of bone and not powder.
I’ll try to drill in about 1.5 inches, collect the powder, decalcify and then hit it with proK to kill any lingering proteins in there. I’m going to make up fresh reagents this week. I want everything to be right. Will likely test for nuclear and mt DNA fragments next weekend.
It’s all about the specimens on your end, and you really came through with this one…. I won’t hurt it too bad, I promise.
Best,
AC
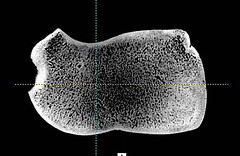 Andy found all kinds of DNA in the first samples we sent–horse, pig, cow, deer, human, etc. No sloth DNA though. The bones were small and broken–just fragments. They had probably been soaking up rural Iowa DNA for years. Now Andy has the thickest, densest, unbroken bone we have. The ICLIC people did a CT scan and it looks perfect inside too.
Andy found all kinds of DNA in the first samples we sent–horse, pig, cow, deer, human, etc. No sloth DNA though. The bones were small and broken–just fragments. They had probably been soaking up rural Iowa DNA for years. Now Andy has the thickest, densest, unbroken bone we have. The ICLIC people did a CT scan and it looks perfect inside too.
Cloning DNA has become routine–the stuff of 7th grade science classes, but contamination remains the biggest challenge of ancient DNA study. DNA is everywhere—people are constantly shedding it in dry skin cells, etc., not to mention what they carry around belonging to family members, pets, etc., and the microbes that live everywhere. Chemical changes, water and warm temps begin breaking down DNA after death, while a host of decay organisms feed on the organic molecules and each other. Retrieving verifiable ancient sequences from this rich DNA stew after thousands of years is a major achievement.  Many an announcement of an ancient DNA discovery has been withdrawn when later analysis showed it merely to be the result of contamination. For that reason special labs like the Ancient DNA Centre have been created with personnel trained in techniques designed to reduce the risk of errors. No one has ever sequenced any part of the Megalonyx genome, but if anyone can do it, Andy can. . . . Dave
Many an announcement of an ancient DNA discovery has been withdrawn when later analysis showed it merely to be the result of contamination. For that reason special labs like the Ancient DNA Centre have been created with personnel trained in techniques designed to reduce the risk of errors. No one has ever sequenced any part of the Megalonyx genome, but if anyone can do it, Andy can. . . . Dave


Very cool!
Hi Pete, Very cool indeed. Andy sent some photos of the lab. I’ll get those up tomorrow. Follow the link to Andy’s lab–they are doing some amazing things–practically the stuff of Jurassic Park.